Our Research
Research Areas
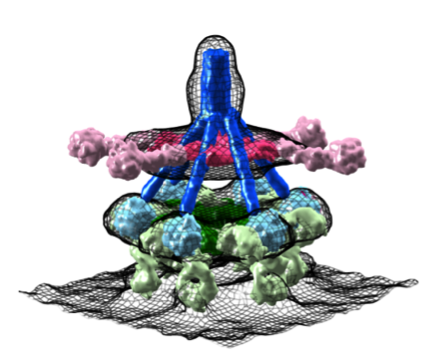
Complement
The complement system is an evolutionary ancient immune defense mechanism, present in vertebrates and invertebrates. It forms a first line of defense in plasma and fluids surrounding tissues and enables the host to recognize and clear invading pathogens (bacteria and viruses) and altered host cells while protecting healthy host cells and tissues. In humans this system is formed by more than 30 large modular plasma proteins and cell surface receptors.
We study the complement system using a variety of biophysical techniques including X-ray Crystallography and Cryo-Electron Microscopy.
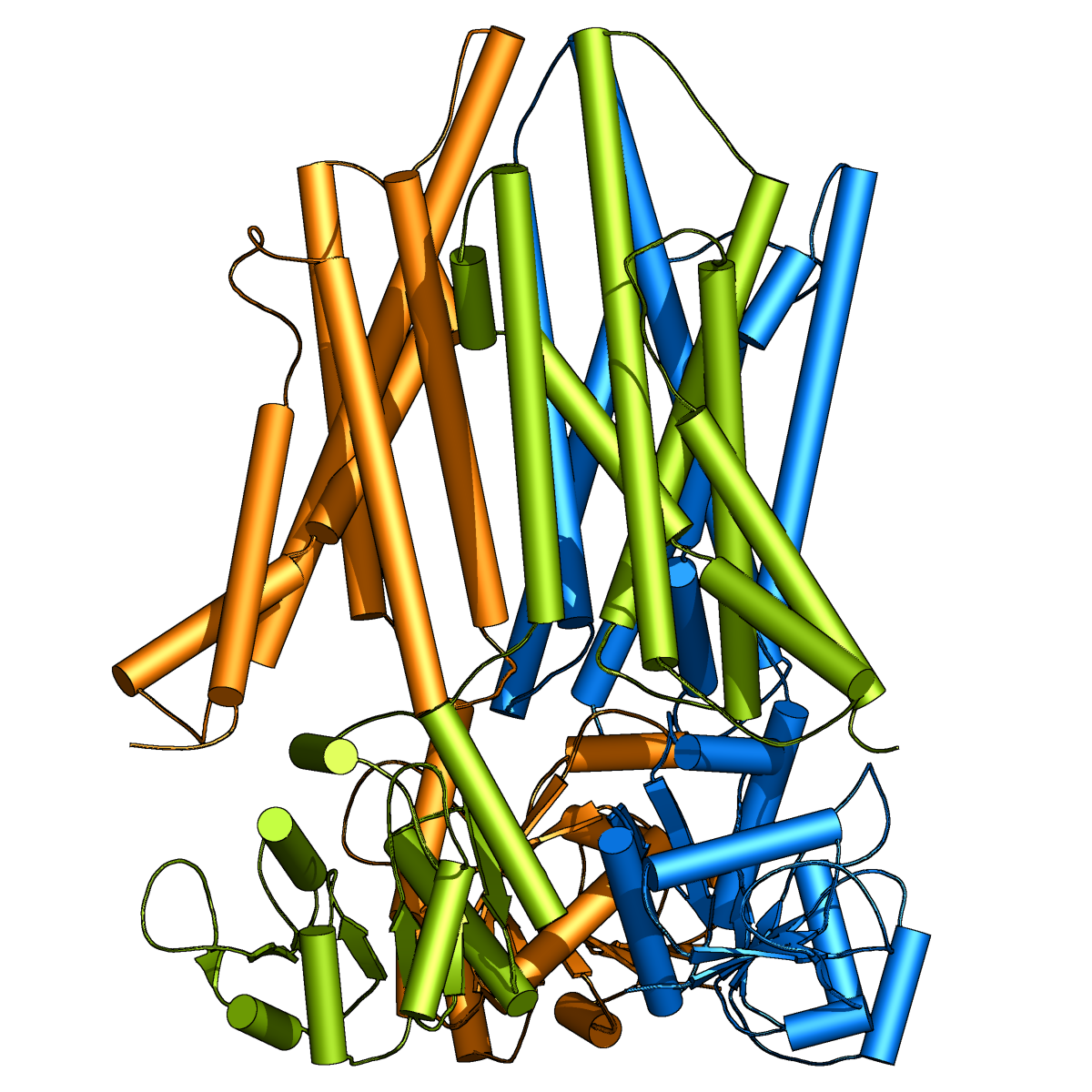
Membrane Proteins
Membrane proteins comprise 20-30% of proteins in the genome, but are targeted by over 50% of modern pharmaceuticals.
We study a number of membrane proteins using a variety of biophysical techniques including X-ray Crystallography and Cryo-Electron Microscopy.
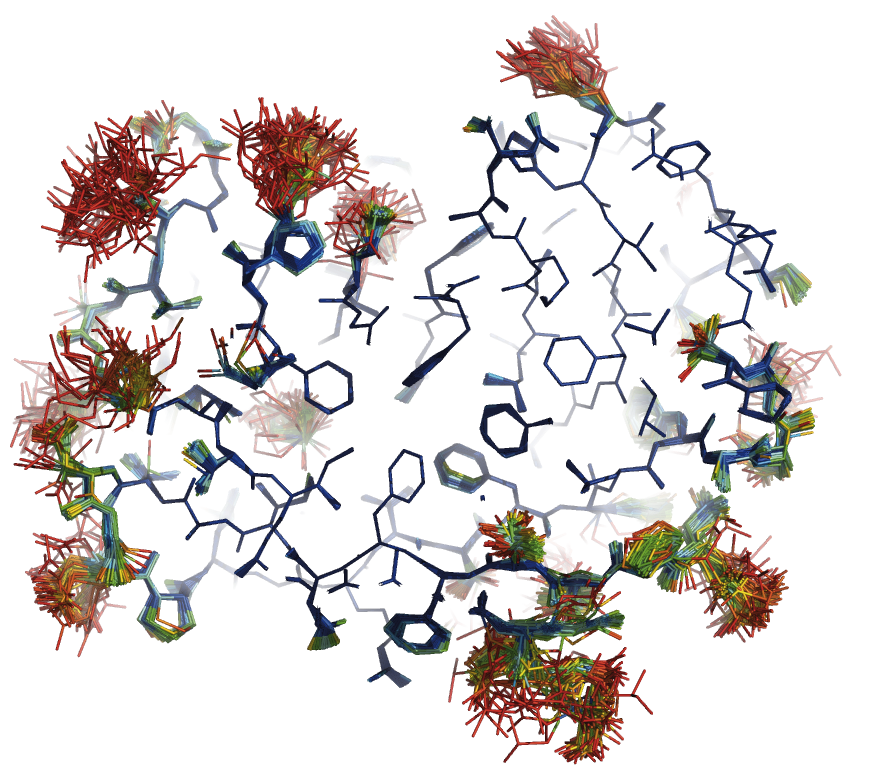
Crystallographic Methods
We have worked on a variety of crystallographic methods for determining molecular structures or proteins with crystallography. These methods include:
Multi-dataset crystallographic methods, utilising multiple crystallographic datasets during structure solution
Ensemble Refinement, a method for combining Molecular Dynamics simulations with experimental diffraction data
Conditional Optimisation, a novel method for refinement, automated model building and ab initio phasing







































